Signatera™ Indications
Signatera™ can be used for a variety of cancer types. To learn more about how, please select your role and a cancer type.
Why circulating tumor DNA (ctDNA) for molecular residual disease assay (MRD) assessment?
- ctDNA is a powerful tool that can be measured to assess the absence or presence of MRD
- Signatera™ is a highly sensitive and personalized MRD assay using ctDNA and is custom designed for each patient to help identify relapse earlier than standard of care tools
Clinical applications of ctDNA testing for MRD assessment
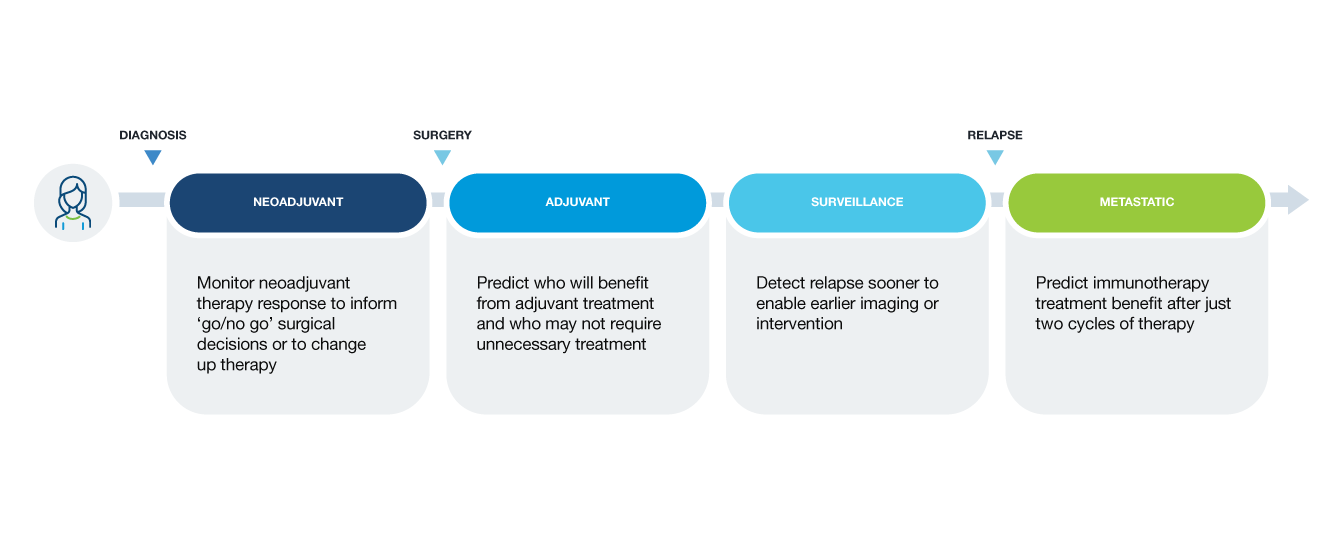

Not all MRD tests are created equal
Signatera™ is the most comprehensive MRD test available. Review our peer-reviewed publications here.
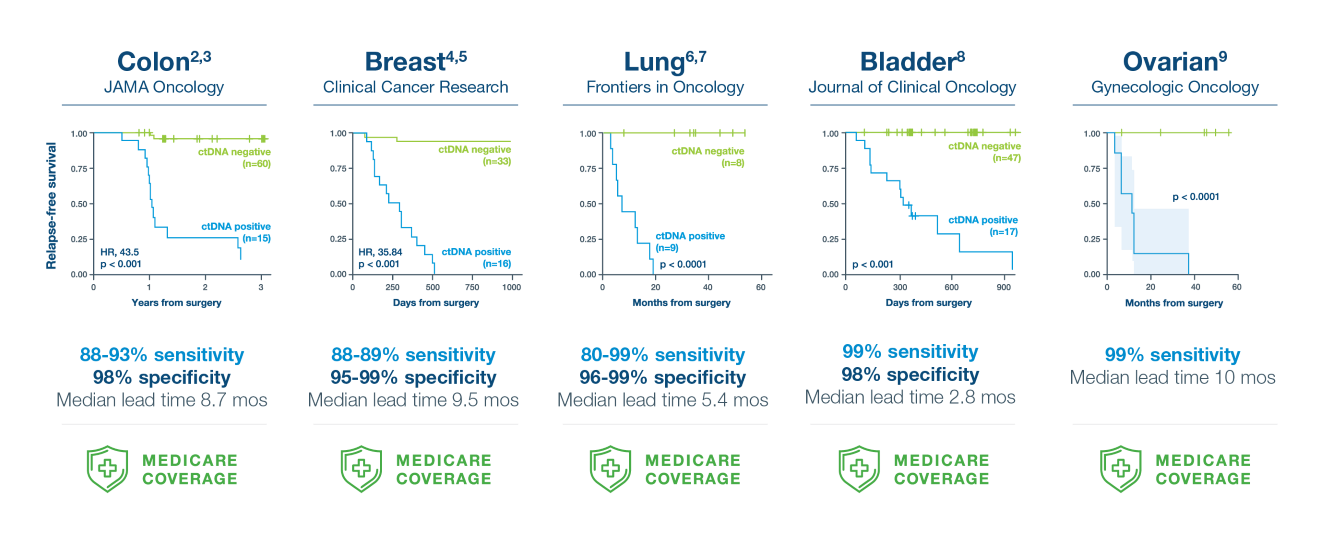
A positive Signatera™ result predicts relapse with overall positive predictive value more than 98%2-9
In clinical studies, Signatera showed high performance across multiple solid tumors
Signatera™ is the first tumor-specific assay for truly individualized cancer care
Personalized design for every patient
- Custom-built assay—based on the unique mutation signature of each patient’s tumor—identifies and tracks tumor mutations at the source
- Once a personalized assay is designed, a patient’s blood can be used to accurately monitor for the presence or absence of the disease over time
How Signatera™ Works: a personalized and tumor informed approach to MRD surveillance

Personalized, tumor-informed assay
One-time, primary tissue sample and matched normal sample is required for whole exome or whole genome sequencing and personalized test design.
Ultrasensitive ctDNA detection
Signatera™ is designed to detect ctDNA of somatic and truncal variants to optimize sensitivity. Tumor-informed method enables filtering of CHIP mutations to decrease false positive rates.
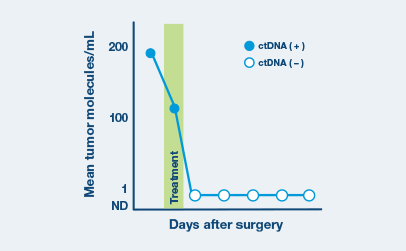
Optimized for longitudinal monitoring
Once the patient’s personalized test has been designed, only a blood sample is needed each subsequent time.
Covered by Medicare for multiple solid tumor indications
Is Signatera™ right for you?
We’re here to help you find out
References
1Natera Data on File as of July 1st, 2025.
2Reinert T, Henriksen TV, Christensen E, et al. JAMA Oncol. 2019.
3Kotani D. et al. Nature Medicine v29 Issue 1 Jan 2023
4Coombes RC, Page K, Salari R, et al. Clin Cancer Res. 2019;25(14):4255-4263
5Shaw et al. JCO Precis Oncol 8, e2300456(2024). DOI:10.1200/PO.23.00456.
6Lebow, E. et al. Front. Oncol. 2023,13:1253629.
7Martin T, Dinerman A, Sudhaman S, et al. J Thorac Cardiovasc Surg. 2024.
8Christensen E, Birkenkamp-Demtroder K, Sethi H, et al. J Clin Oncol. 2019;37(18):1547-1557.
9Hou JY, et al. Gynecologic Oncology. 2022; 167: 334-341.







